Technical Note 12 – Best Practice for Nucleic Acid Quantifications
Nucleic Acid quantification is an important step for many different life science protocols including those implemented for NGS, qPCR and several others. The NanoPhotometer® offers preprogrammed Nucleic Acid methods with options to measure all types of nucleic acids including dsDNA, ssDNA, RNA, miRNA, and Oligos, as well as the option to enter sequences for miRNAs and Oligos as a custom defined extinction coefficient for other types of nucleic acids. It is also possible to measure fluorescent labeled nucleic acids, e.g. as a quality control step during micro array workflows, via the absorbance maxima of the dye. Dye concentration and the frequency of incorporation are automatically calculated and displayed.

Figure 1: The Pedestal and mirror are being cleaned with a lint-free tissue
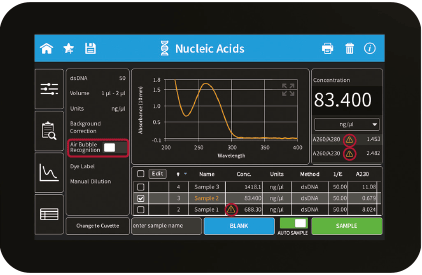
Figure 2: Result screen with Measurement Bar, Air Bubble Recognition and Sample Control
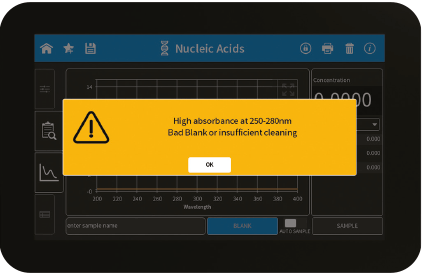
Figure 3: Bad Blank warning
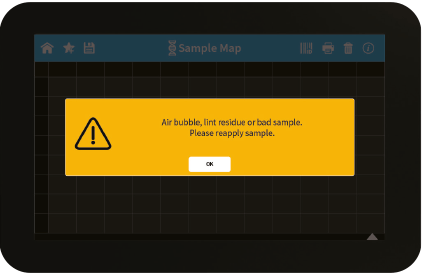
Figure 2: Air Bubble/Lint residue warning.
Nucleic Acid Quantification Steps
To obtain optimal readings we suggest the following steps:
- Clean the measurement head (both bottom and top/mirror) using a lint free tissue (avoid vigorous polishing to prevent lint deposits on the quartz window)
- If the pedestal is dirty, apply 5 µl of water onto pedestal and close the lid to soak the spot for at least one minute and wipe pedestal and mirror thoroughly afterwards
- Blank with the same buffer used to elute/dissolve the sample
Recommended buffer for nucleic acids: 10 mM Tris-HCl buffer, pH 8.0 - Use low-binding tubes and pipette tips for samples with low concentrations
- Measure the blanking solution as a sample to identify a bad blank
- Homogenize the sample using the built-in vortexer or by pipetting up/down (gDNA)
- Use the illuminated pedestal to confirm the sample is properly positioned with no visible bubbles
- Enable “Air Bubble Recognition” if using buffers containing detergents such as Tween or Triton
- Measure replicates if sample concentration is <10 ng/µl, as greater % error is expected
- Check 260/230, 260/280 ratios and full spectral graph data for contaminants. Sample Control will flag bad samples.
- Properly clean pedestal after each use
Nucleic Acid Quantification Important Warning Messages:
| Message Text | Explanation/Solution |
| Air bubble, lint residue or bad sample. Please reapply sample. | Air bubble recognition is on and has detected either an air bubble, lint residue or a bad sample like e.g. a turbid sample. Check sample, clean the sample window on pedestal and mirror in the lid arm thoroughly and reapply sample carefully. To avoid air bubbles apply sample by reverse pipetting. |
| High absorbance at 250 – 280 nm, 280 – 340 nm, 340 – 400 nm, 400 – 475 nm, 475 – 550 nm, 550 – 625 nm, 625 – 700 nm. Bad Blank or insufficient cleaning. | The warning message is shown when a blank measurement (NanoVolume) detects a significant absorbance a potential area of interest. The warning message shows the wavelength range where the absorbance is appearing. Two things can cause high absorbance in a blank measurement: either the blank solution/buffer has an absorbance in this wavelength range or the sample window on pedestal and the mirror in the lid arm was not cleaned properly after the last reading. Clean the sample window on pedestal and the mirror in the lid arm thoroughly and check the blank solution for absorbance (Use water for blank and measure the blank buffer as sample). |
| Lint residue or bad sample. Please reapply sample. | Displayed when Air Bubble Recognition is off. Software has detected either a lint residue or a bad sample like e.g. a turbid sample. Check sample, clean the sample window on pedestal and mirror in the lid arm thoroughly and reapply sample carefully. |
| Maximum absorbance level at specified wavelength reached. Calculations may lead to low/wrong results. | The concentration of the sample used is too high and exceeds the specified absorbance range. Maximum absorbance is 330 A (10 mm path) for NanoVolume and 2.65 A for cuvette measurements. Dilute sample and measure again. |
| Maximum level exceeded. | This message is shown when for example, too many dyes or wavelength are added with the Add + button. It is possible to add up to 20 dyes or wavelengths. |
| Sample concentration too low for 0.3 µl – change volume and parameter settings to 1 µl | The measurement parameters for 0.3 µl NanoVolume samples read only the 0.07 mm path length (dilution 140). The utilized sample concentration is too low. Please use 1 µl of the sample and change the volume setting to 1 – 2 µl. Minimum concentrations for the 0.07 mm path length for dsDNA 420 ng/µl and for BSA 12.6 mg/ml |
For further questions or if assistance is needed, please contact your local Implen Support Team at [email protected] or [email protected].